Biotechnology of the Interaction of Microorganisms with Legumes and Other Plants of Agricultural Interest
Francisco Javier López Baena
Profesor Titular
Francisco Pérez Montaño (IP)
Profesor Contratado Doctor
Departamento de Microbiología, Facultad de Biología, Universidad de Sevilla (BIO169)
José Manuel Borrero de Acuña
Investigador Postdoctoral (Emergia, Junta de Andalucía).
TELÉFONO
+34 954 557 116
Patricia Bernal Guzmán
Investigadora Postdoctoral Ramón y Cajal
Other group members
Research lines
- Regulation of the production of bacterial molecular signals involved in the rhizobium-legume symbiotic interaction.
- Structural determination of molecular signals involved in rhizobium-legume symbiosis (Nod factors, bacterial surface polysaccharides, flavonoids).
- Determination of the non-coding RNome of Sinorhizobium fredii HH103 and Rhizobium tropici CIAT899 and study of their role in symbiosis with their legume hosts.
- Bacterial secretion systems important in bacteria-plant and bacteria-bacteria relationships: T3SS and T6SS
- Rhizobial effectors of T3SS involved in symbiotic compatibility with wild and improved soybean.
- Role of rhizobial membrane vesicles in symbiosis with legumes.
- Engineering of extracellular membrane vesicles of rhizobial bacteria for the development of biopesticides and plant growth promoting agents.
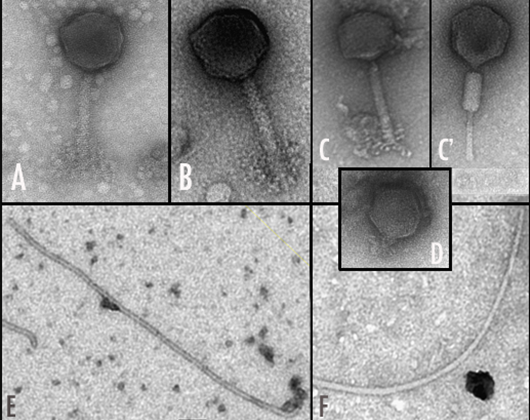
- Isolation and characterisation of rhizobial bacteriophages.
- Molecular mechanisms controlling the growth, colonisation or infection of phytopathogenic bacteria in crops of agricultural interest.
Representative publications
- Acosta-Jurado S, Alías-Villegas C, Navarro-Gómez P, Almozara A, Rodríguez-Carvajal MA, Medina C, Vinardell JM (2020) Sinorhizobium fredii HH103 syrM inactivation affects the expression of a large number of genes, impairs nodulation with soybean, and extends the host-range to Lotus japonicus. Environ Microbiol 22: 1104-1124.
- Cubo MT, Alías-Villegas C, Balsanelli E, Mesa D, de Souza E, Espuny MR (2020) Diversity of Sinorhizobium (Ensifer) meliloti bacteriophages in the rhizosphere of Medicago marina: myoviruses, filamentous and N4-like podovirus. Front Microbiol 11: 22 (doi: 3389/ fmicb.2020.00022).
- Acosta-Jurado S, Alías-Villegas C, Almozara A, Espuny MR, Vinardell, JM, Pérez-Montaño F (2020) Deciphering the symbiotic significance of quorum sensing systems of Sinorhizobium fredii Microorganisms 8: 68 (doi: 10.3390/microorganisms 8010068).
- Di Lorenzo F, Speciale I, Silipo A, Alías-Villegas C, Acosta-Jurado S, Rodríguez-Carvajal MÁ, Dardanelli MS, Palmigiano A, Garozzo D, Ruiz-Sainz JE, Molinaro A, Vinardell JM. (2020) Structure of the unusual Sinorhizobium fredii HH103 lipopolysaccharide and its role in symbiosis. J Biol Chem 295: 10969-10987.
- Jiménez-Guerrero I, Acosta-Jurado S, Medina C, Ollero FJ, Alias-Villegas C, Vinardell JM, Pérez-Montaño F, López-Baena FJ (2020) The Sinorhizobium fredii HH103 Type III secretion system effector NopC blocks nodulation with Lotus japonicus J Exp Bot 71: 6043-6056.
- Jiménez-Guerrero I, Pérez-Montaño F, Zdyb A, Beutler M, Werner G, Göttfert M, Ollero FJ, Vinardell JM, López-Baena FJ (2019) GunA of Sinorhizobium (Ensifer) fredii HH103 is a T3SS-secreted cellulase that differentially affects symbiosis with cowpea and soybean. Plant Soil 435: 15-26.
- Acosta-Jurado S, Rodríguez-Navarro DN, Kawaharada Y, Rodríguez-Carvajal MA, Gil-Serrano A, Soria-Díaz ME, Pérez-Montaño F, Fernández-Perea J, Yanbo N, Alias-Villegas C, Jiménez-Guerrero I, Navarro-Gómez P, López-Baena FJ, Kelly S, Sandal N, Stougaard J, Ruiz-Sainz JE, Vinardell JM (2019) Sinorhizobium fredii HH103 nolR and nodD2 mutants gain capacity for infection thread invasion of Lotus japonicus Gifu and Lotus burttii. Environ Microbiol 21: 1717-1739.
- Temprano-Vera F, Rodríguez-Navarro DN, Acosta-Jurado S, Perret X, Fossou RK, Navarro-Gómez P, Zhen T, Yu D, An Q, Buendía-Clavería AM, Moreno J, López-Baena FJ, Ruiz-Sainz JE, Vinardell JM (2018) Sinorhizobium fredii strains HH103 and NGR234 form nitrogen fixing nodules with diverse wild soybeans (Glycine soja) from Central China but are ineffective on Northern China accessions. Front Microbiol 9: 2843 (doi: 10.3389/fmicb2018.02843).
- Acosta-Jurado S, Navarro-Gómez P, Crespo-Rivas JC, Medina C, Murdoch PS, Cuesta-Berrio L, Rodríguez-Carvajal MA, Ruiz-Sainz JE, Vinardell JM (2017) The Sinorhizobium (Ensifer) fredii HH103 rkp-2 region is involved in the biosynthesis of lipopolysaccharide and exopolysaccharide but not in K-antigen polysaccharide production. Plant Soil 417: 415-431.
- Crespo-Rivas JC, Guefrachi I, Mok KC, Villaécija-Aguilar JA, Acosta-Jurado S, Pierre O, Taga ME, Mergaert P, Vinardell JM (2016) Sinorhizobium fredii HH103 bacteroids are not terminally differentiated and show altered O-antigen in nodules of the IRLC legume Glycyrrhiza uralensis. Environ Microbiol 18: 2392-2404.
- Pérez-Montaño F, Jiménez-Guerrero I, Acosta-Jurado S, Navarro-Gómez P, Ollero FJ, Ruiz-Sainz JE, López-Baena FJ, Vinardell JM (2016) A transcriptomic analysis of the effect of genistein on Sinorhizobium fredii HH103 reveals novel rhizobial genes putatively involved in symbiosis. Sci Rep 6: 31592 (doi: 10.1038/srep31592).
- Acosta-Jurado S, Navarro-Gómez P, Murdoch PdelS, Crespo-Rivas JC, Jie S, Cuesta-Berrio L, Ruiz-Sainz JE, Rodríguez-Carvajal MA, Vinardell JM (2016) Exopolysaccharide production by Sinorhizobium fredii HH103 is repressed by genistein in a NodD1-dependent manner. PLoS One 11:
- Acosta-Jurado S, Alias-Villegas C, Navarro-Gómez P, Zehner S, Murdoch PD, Rodríguez-Carvajal MA, Soto MJ, Ollero FJ, Ruiz-Sainz JE, Göttfert M, Vinardell JM (2016) The Sinorhizobium fredii HH103 MucR1 Global Regulator Is Connected With the nod Regulon and Is Required for Efficient Symbiosis With Lotus burttii and Glycine max Williams. Mol Plant Microbe Interact 29: 700-712.
- Acosta-Jurado S, Rodríguez-Navarro DN, Kawaharada Y, Perea JF, Gil-Serrano A, Jin H, An Q, Rodríguez-Carvajal MA, Andersen SU, Sandal N, Stougaard J, Vinardell JM, Ruiz-Sainz JE (2016) Sinorhizobium fredii HH103 invades Lotus burttii by crack entry in a Nod factor-and surface polysaccharide-dependent manner. Mol Plant Microbe Interact 29: 925-937.
- Vinardell JM, Acosta-Jurado S, Göttfert M, Zehner S, Becker A, Baena-Ropero I, Blom J, Crespo-Rivas JC, Goesmann A, Jaenicke S, Krol E, McIntosh M, Margaret I, Pérez-Montaño F, Schneiker-Bekel S, Serrania J, Szczepanowski R, Buendia-Claveria AM, Lloret J, Bonilla I, Pühler A, Ruiz-Sainz JE, Weidner S (2015) The Sinorhizobium fredii HH103 genome: a comparative analysis with fredii strains differing in their symbiotic behaviour with soybean. Mol Plant Microbe Interact 28: 811-824.
Grants
- Estudios de mecanismos de regulación alternativos para la síntesis de señales simbióticas en rizobios. PID2022-141156OB-I00. 01/09/2023 – 31/08/2026
- ¿Antibiosis o simbiosis? Caracterizando el sistema de secreción de tipo VI de Sinorhizobium fredii usda257. PID2020-118279RA-I00. 01/09/2021 – 31/08/2024
- Mejorando el diálogo simbiótico a través de vesículas de membrana rizobianas. PID2021-122395OA-I00. 01/09/2022 – 31/08/2026
- Engineering extracellular membrane vesicles from rhizospheric bacteria for the development of biopesticides and plant-growth promoting agents. PROYEXCEL_00450. 02/12/2022 – 31/12/2025.
- Desarrollo de pesticidas biológicos basados en vesículas de membrana como alternativa sostenible a los pesticidas químicos altamente contaminantes. TED2021-130357B-I00. 01/12/2022 – 30/11/2024
- Señalización sistémica en la simbiosis rizobio-leguminosa y nutrición nitrogenada. Efectos sobre la productividad vegetal. PID2021-122353OB-I00. 01/09/2022 – 31/08/2025
Relevant methods
| SUPPORT | METHODOLOGY |
| 1. Thermo scientific liquid chromatography system. | Determinación de factores de nodulación. |
| 2. Nanosight ns300 marvern panalitycal nanoparticle analyzer. | Aislamiento de vesículas extracelulares de membrana. |
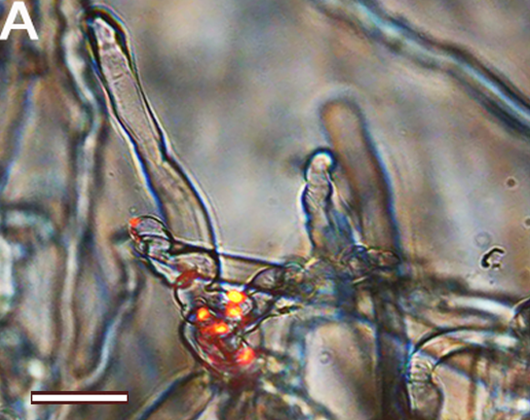
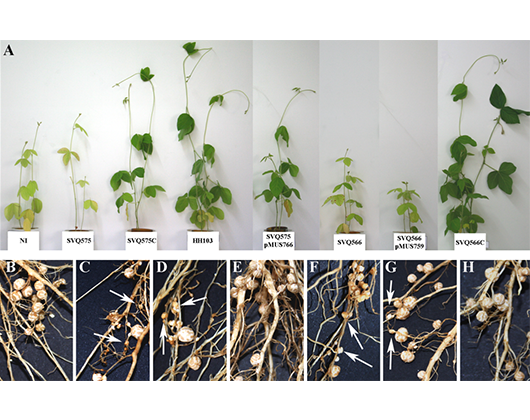
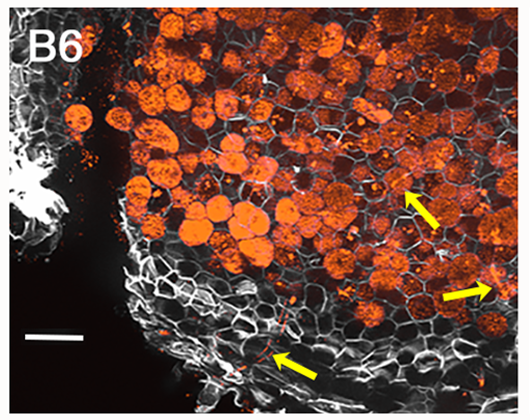
| 3. Zeiss fluorescence apotome.2 microscope. | Visualización del progreso de la infección: colonización, tubos de infección y nódulos. Localización de proteínas e interacción entre proteínas en hojas de tabaco. |
| 4. Cromatógrafo de gases. | Cuantificación de la fijación de nitrógeno por ARA (acetylene reduction assay) |
Collaborations with other national and international research groups
- David S. Guttman – University of Toronto, Canadá
- Dieter Jahn – Technischen Universität Carolo-Wilhelmina Zu Braunschweig, Alemania
- Sussane Sievers – Universität Greifswald, Alemania
- Federico Martinelli – Università Degli Studi di Firenze, Italia
- Ignacio Poblete Castro – Universidad de Santiago de Chile
- Saul Burdman – The Hebrew University of Jerusalem, Israel
- Maria Camacho Martinéz Vara Del Rey – IFAPA, España
- Alessio Mengoni – Università Degli Studi di Firenze, Italia
- José Ignacio Jiménez Zurdo – Estación Experimental del Zaidín, España
- María José Soto Misfutt – Estación Experimental del Zaidín, España
- Myriam Charpentier – John Innes Centre, Reino Unido
- Juan Sanjuan Pinilla – Estación Experimental del Zaidín, España
- Jens Stougaard – Aarhus Universitet, Dinamarca
- Flaviana Di Lorenzo – Università Degli Studi Di Napoli, Italia
- Mariangela Hungria – EMBRAPA Soja, Brasil
- Thomas Martin – Universidade Federal de Santa Maria, Brasil
- Alain Filloux – Imperial College of London, Reino Unido
- Despoina Mavridou – University of Texas at Austin, EEUU
- George diCenzo – Queen’s University, Kingston, Ontario, Canadá
- Chang-Fu Tian – China Agricultural University, Beijing, China
- Zhen-Tao – Heilongjiang Academy of Sciences, Harbin, China
- Peter Kaló – Institute of Plant Biology, HUN-REN Biological Research Centre, Szeged, Hungría